Small Language Models Meet the Clinic: Benchmarking 7B-9B Systems on Clinical Information Extraction
What 65,000 predictions on real physician notes reveal about SLM readiness

A hospital wants to automate the extraction of structured information from discharge letters: strip patient identifiers before sharing data for research, flag bacterial infections and their negation status, classify treatment outcomes, build structured patient profiles. The promise of general-purpose small language models (SLMs) replacing task-specific pipelines is appealing: a single 7-9 billion-parameter model, running on a single GPU, handling all four tasks out of the box.
But does it hold?
We ran seven systems across four clinical information extraction tasks, generating 65,065 predictions on real physician-written notes. The headline finding inverts the intuition that bigger means better: a 200M-parameter encoder GLiNER2 outperforms every 7-9B LLM by a factor of three on aggregate zero-shot performance. More surprising still, the bottleneck is not clinical understanding. The models often know the right answer but cannot reliably produce a valid structured output. Schema conformity is the binding constraint.
This report presents the full results, dissects the failure modes, and offers actionable guidance for clinicians, hospital IT teams and AI engineers considering SLMs for clinical IE. The evaluation corpus is PARHAF, a recently released collection of thousands of realistic clinical notes written by practicing physicians in French. While the data is in French, the core findings about architectural fit, output discipline, and few-shot behavior may apply broadly to clinical IE.
All code and configurations are open-source at github.com/lounesmechouek/parhaf-clibench
Predictions, errors and insights can be viewed in a complementary interface at parhaf-clibench.streamlit.app
What We Tested and Why It Matters
The PARHAF Dataset
Most clinical NLP benchmarks rely on synthetic notes or heavily curated subsets. PARHAF breaks this pattern. Released in late March 2026 by the Health Data Hub, it is a large-scale corpus of realistic French clinical notes written by practicing physicians. The notes cover 21 medical specialties and reflect the messy, abbreviated, jargon-dense style of real hospital documentation.
Four annotated subsets define our evaluation tasks:
| Subset | Documents | Annotated records | Source |
|---|---|---|---|
| PARHAF-pseudo-annotated | 509 | ~7,000 identifying spans | HuggingFace |
| PARHAF-infectiology-annotated | 134 | ~3,600 clinical mentions | HuggingFace |
| PARHAF-response_to_treatment-annotated | 108 | 108 treatment outcomes | HuggingFace |
| PARHAF (structured scenario) | 4,254 | ~13,600 structured fields | HuggingFace |
Every dataset is pinned to a specific Git revision for reproducibility. The total evaluation footprint: 5,005 documents, 65,065 predictions, 25,880 error records.
Four Clinical IE Tasks
Each task transforms a clinical discharge note into a list of structured records. The tasks span a range of difficulty, from character-level span extraction to document-level classification.
| Task | What it extracts | Official metric | Why it is hard |
|---|---|---|---|
| Pseudonymization | Patient identifiers (names, dates, addresses, phone numbers) with exact character offsets | Micro-F1 on (start, end) byte pairs |
Requires byte-level positional accuracy from a text generator |
| Infectiology | Bacteria, infections, and anatomical sites with negation status (Present / Absent / Indeterminate) | Micro-F1 on (text, label, negation) triples |
Negation detection in clinical prose; three-way attribute classification |
| Response to treatment | Treatment outcome classification with justification span | Micro-F1 on (text, label) pairs |
Six-class problem over specialized vocabulary; small corpus (108 docs) |
| Structured scenario | Eight structured fields (name, age, sex, admission/discharge mode, diagnosis, procedure, type of care) | Micro-F1 on (text, label) pairs |
Closest to open-domain QA; 8 distinct extraction targets across 21 specialties |
The pseudonymization task is intentionally the strictest: the evaluation requires exact character offsets (start, end), not just correct entity text. This choice reflects real-world constraints. If spans are not taken into account, the system may mask the wrong segments. For example, the same surface form can appear once as non-sensitive and elsewhere as personal information. Without precise span alignment, both occurrences could be masked indiscriminately. While this over-masking is less critical than failing to mask sensitive data, our design enforces strict span accuracy. This is justified because specialized encoder models such as GLiNER consistently predict correct spans, and we want to explicitly measure whether generators can match this capability.
The structured scenario task sits at the opposite end: it asks the model to extract high-level patient attributes from free text, more like a reading comprehension exercise than a sequence labeling task.
Clinical discharge note
========================
"Le patient M. Jean Dupont, 67 ans, a ete admis le 12/03/2025
pour pneumonie bacterienne a Klebsiella pneumoniae..."
| | | |
v v v v
PSEUDONYMIZATION INFECTIOLOGY RESPONSE TO TX STRUCTURED SCENARIO
----------------- --------------- --------------- --------------------
"Jean Dupont" Klebsiella Partial response name: Jean Dupont
start=X, end=Y pneumoniae (justification: age: 67
label: LAST_NAME label: Bacterie "amelioration sex: M
"12/03/2025" negation: Present partielle...") admission: urgences
start=W, end=Z diagnosis: pneumonie
label: DATE ...
Each task transforms the same clinical note into a different structured output. Pseudonymization requires exact character offsets; infectiology requires negation attributes; the scenario task extracts eight high-level fields.
The Seven Systems
We evaluate six instruction-tuned LLMs in the 7-9B parameter range and one encoder-based baseline:
| System | Parameters | Architecture | Context window | Notes |
|---|---|---|---|---|
| GLiNER2-multi | ~200M | Encoder (span extraction) | 512 tokens | Zero-shot NER; purpose-built for entity extraction |
| Gemma2-9B-it | 9B | Decoder (LLM) | 8,192 tokens | Google, instruction-tuned |
| Ministral-8B-Instruct | 8B | Decoder (LLM) | 32,768 tokens | Mistral AI, instruction-tuned |
| Qwen2.5-7B-Instruct | 7B | Decoder (LLM) | 32,768 tokens | Alibaba, instruction-tuned |
| Llama-3.1-8B-Instruct | 8B | Decoder (LLM) | 32,768 tokens | Meta, instruction-tuned |
| Aya-Expanse-8B | 8B | Decoder (LLM) | 8,192 tokens | Cohere, 23-language multilingual |
| Lucie-7B-Instruct | 7B | Decoder (LLM) | 4,096 tokens | OpenLLM-France, French-focused |
The design space is intentional: five general-purpose multilingual LLMs, one French-specialized model (Lucie), and one non-autoregressive encoder that represents the current state of the art for zero-shot span extraction. All LLMs are served through vLLM with structured-output JSON schema enforcement via the xgrammar backend. Inference runs on a single NVIDIA RTX A6000 GPU with identical parameters across all models: temperature 0, top-p 1, seed 42, max output tokens 512.
GLiNER2 runs its own runtime with post-inference schema compliance and heuristic negation detection.
Evaluation Protocol
Two evaluation tracks:
- Zero-shot: The model receives a task instruction, a canonical JSON schema describing the expected output format, and the source clinical note. No examples.
- Few-shot fixed: The same prompt, augmented with three frozen, synthetically generated demonstration examples. The bank is identical across all models and held constant throughout the evaluation.
All prompts are written in French (matching the corpus language), versioned by SHA-256 hash, and rendered from Jinja2 templates.
Scoring:
- Per-task metric: Micro-F1 computed over task-specific elementary units (character spans for pseudonymization, text-label-negation triples for infectiology, text-label pairs for the other two tasks).
- Global score: Equal-weight arithmetic mean of the four task F1 scores. This deliberate choice prevents the scenario task (30x more documents) from dominating the aggregate.
- Uncertainty quantification: Document-level non-parametric bootstrap with B = 1,000 replications, yielding 95% percentile confidence intervals (2.5th to 97.5th percentile). Paired bootstrap for head-to-head model comparisons.
- Robustness metrics: Schema conformity rate, empty output rate, raw JSON validity rate, median latency, and throughput for every (model, track, task) cell.
Scoring audit: An independent re-scoring pipeline reloads all 65,065 predictions from raw JSONL files, re-parses them through the canonical Pydantic schema, reloads gold examples from the offline cache, and re-invokes the scoring modules. All 52 scoring cells reproduce the shipped values to six decimal places.
Clinical note Prompt template Model inference Output parsing
+--------------+ +----------------+ +----------------+ +----------------+
| Discharge | ---> | Jinja2 render | ---> | vLLM / GLiNER2 | --> | JSON parse + |
| letter | | (task + track | | (temp=0, | | schema valid. |
| (French) | | + schema) | | seed=42) | | + offset align |
+--------------+ +----------------+ +----------------+ +----------------+
|
v
Bootstrap CI Micro-F1 scoring Canonical records
+----------------+ +----------------+ +----------------+
| B=1000 doc- | <--- | TP/FP/FN per | <-- | {label, text, |
| level replic. | | task-specific | | start, end, |
| 95% percentile | | elementary | | attributes} |
+----------------+ | units | +----------------+
+----------------+
The evaluation pipeline: from raw clinical notes through prompt rendering, model inference, output parsing, task-specific scoring, and bootstrap confidence intervals.
The Global Picture
Encoders Still Win
The global leaderboard tells a clear story. GLiNER2, with roughly 200 million parameters and no task-specific training, achieves a global F1 of 0.206 on zero-shot. The best LLM on zero-shot is Ministral-8B at 0.074, nearly three times lower. Even the best few-shot LLM (Ministral at 0.166) does not reach GLiNER2’s zero-shot baseline.
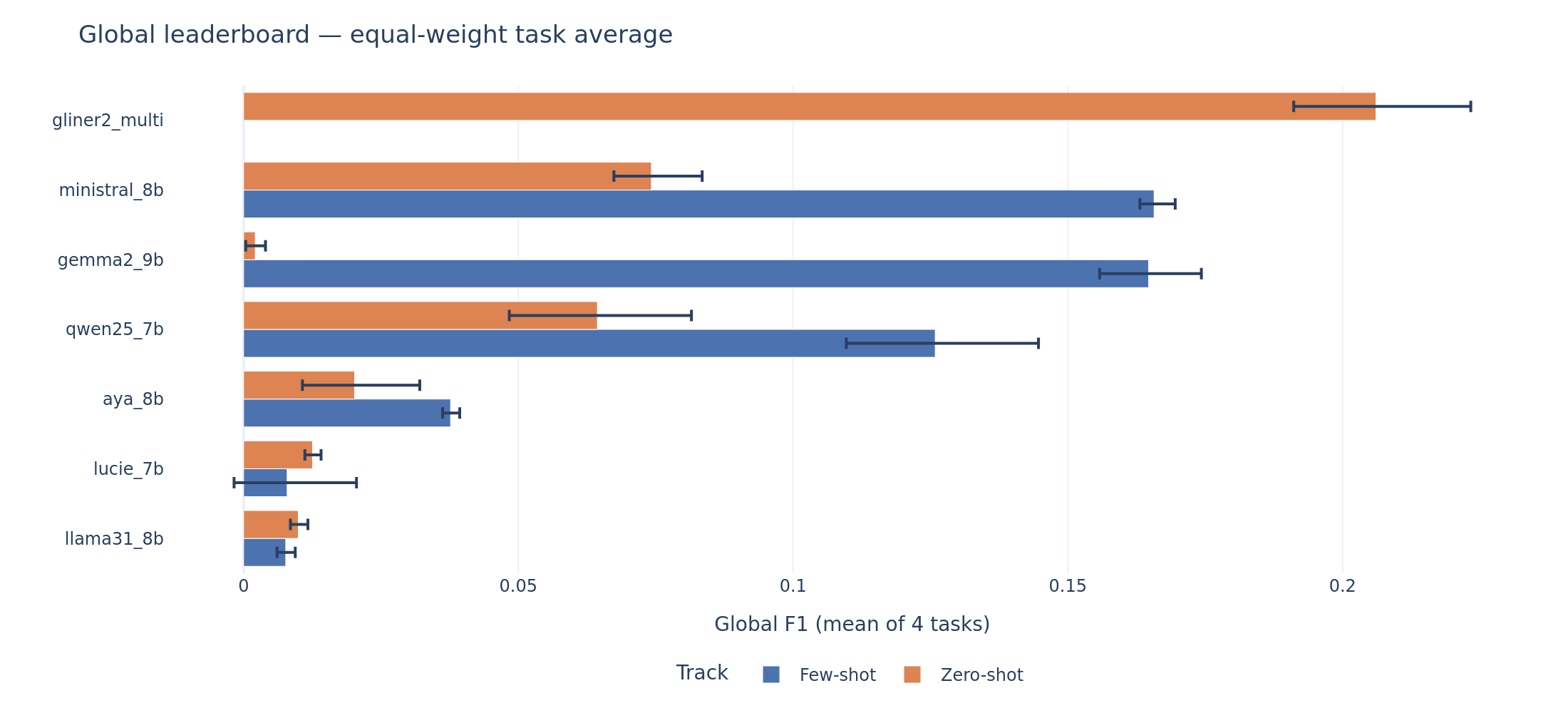
Global leaderboard. Each bar shows the equal-weight mean F1 across four tasks with 95% bootstrap confidence intervals. GLiNER2 zero-shot (top) sets the ceiling that no LLM reaches, even with few-shot demonstrations.
No system on any task reaches the performance levels that the clinical NLP literature and regulatory frameworks associate with safe autonomous deployment. For context: the i2b2/2014 de-identification shared task, the closest English-language benchmark to our pseudonymization task, saw winning systems achieve F1 above 0.95. The EU AI Act classifies clinical decision support as high-risk, requiring demonstrated accuracy and robustness. The FDA’s guidance on AI/ML-based Software as a Medical Device (SaMD) expects performance validation against clinically meaningful thresholds.
We adopt two pragmatic reference points informed by these precedents: F1 >= 0.70 as a minimum for supervised deployment (consistent with operational NER thresholds in prior clinical evaluation campaigns) and F1 >= 0.40 as a minimum for human-in-the-loop piloting. These are not universal cutoffs. The appropriate threshold depends on the downstream risk: pseudonymization for GDPR compliance demands near-perfect recall, while scenario extraction for pre-annotation tolerates more noise. Under either threshold, no system in this benchmark qualifies for autonomous clinical use on any task.
Task-by-Task Breakdown
The aggregate hides important task-level dynamics. The per-task forest plots reveal where each system excels and where it collapses.
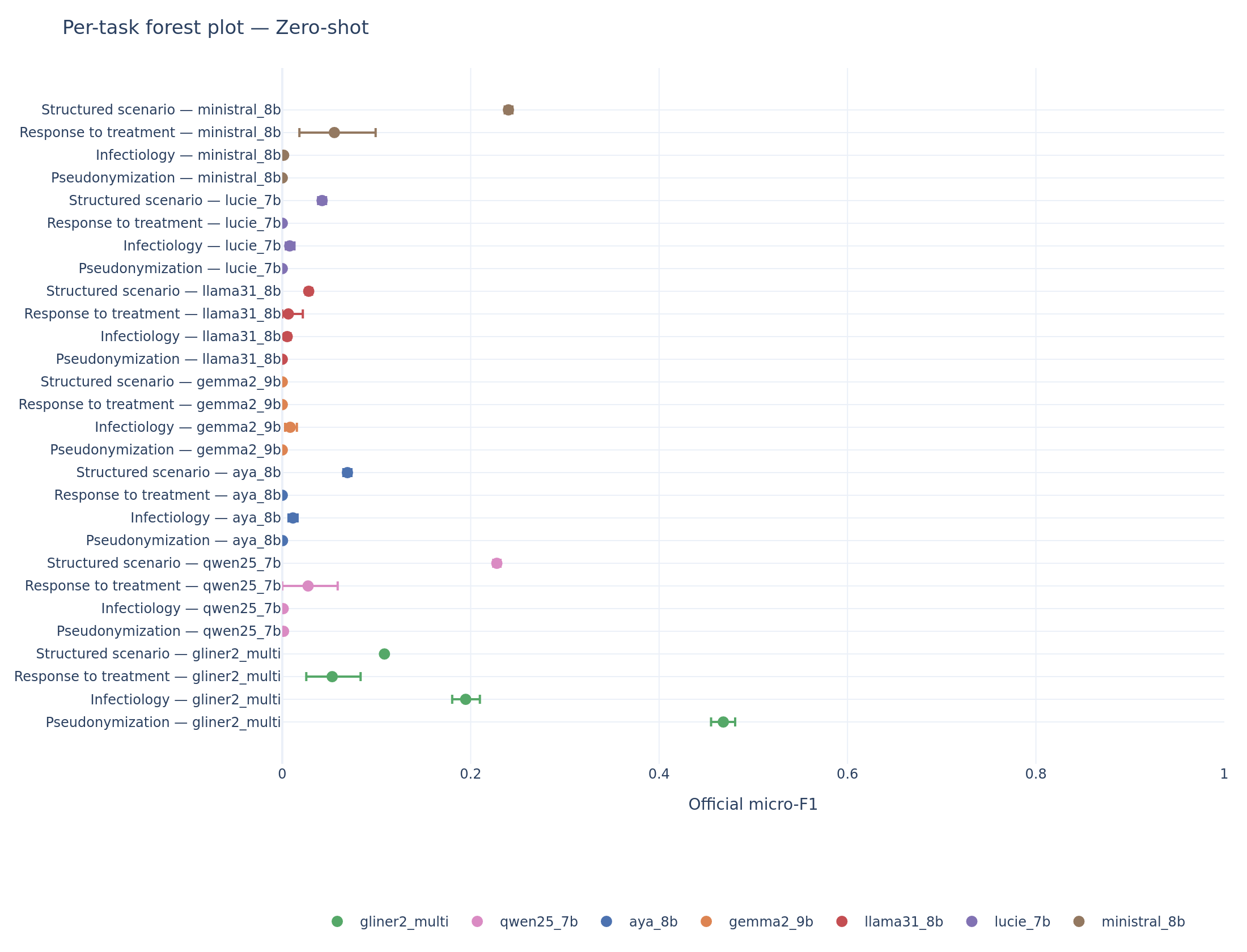
Per-task forest plot (zero-shot). GLiNER2 dominates pseudonymization (F1 = 0.47) and infectiology (F1 = 0.19). On scenario extraction, Ministral and Qwen overtake the encoder.
Pseudonymization is nearly unsolved by LLMs. The official metric requires exact (start, end) character offsets. Every LLM scores below F1 = 0.002 on zero-shot, effectively zero. GLiNER2, designed for span extraction, reaches 0.468. The gap is not about entity recognition: when we relax the metric to normalized text matching (ignoring offsets), Ministral scores 0.218 and Qwen scores 0.292 on zero-shot. The models identify the right entities but cannot count characters. This is a fundamental limitation of autoregressive decoding for positional tasks.

Pseudonymization (zero-shot). The visual gap between GLiNER2 and every LLM captures the core architectural mismatch: text generators cannot reliably produce byte-level positional outputs.
Structured scenario extraction is where LLMs compete. On scenario, the task closest to open-domain question answering, Ministral zero-shot (F1 = 0.240) and Qwen zero-shot (F1 = 0.228) both outperform GLiNER2 (F1 = 0.108). GLiNER2’s low precision on scenario (0.060) reflects an architectural mismatch in reverse: the encoder was not designed for multi-field structured extraction from long narratives.
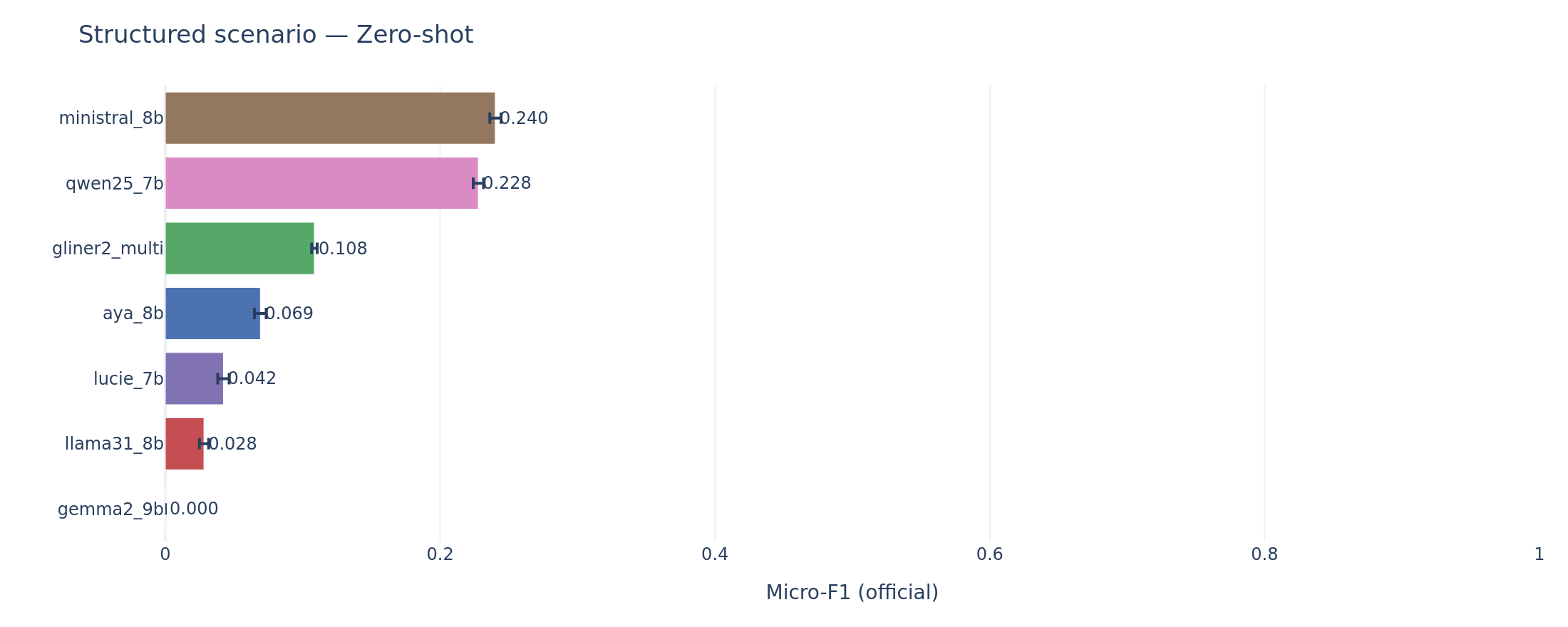
Scenario extraction (zero-shot). The reversal: Ministral and Qwen use their generalist reading comprehension to outperform the span-oriented encoder.
Infectiology follows the encoder-dominant pattern: GLiNER2 at F1 = 0.195 leads all LLMs, with the best zero-shot LLM (Aya) at only 0.011. Negation handling is the key differentiator: detecting “absence of infection” in clinical French requires both mention extraction and attribute classification.
Response to treatment is low across the board. Ministral zero-shot leads at F1 = 0.055, barely above GLiNER2 at 0.053. The small corpus (108 documents) produces wide confidence intervals, making it hard to draw firm conclusions.
The paired bootstrap quantifies these comparisons statistically:
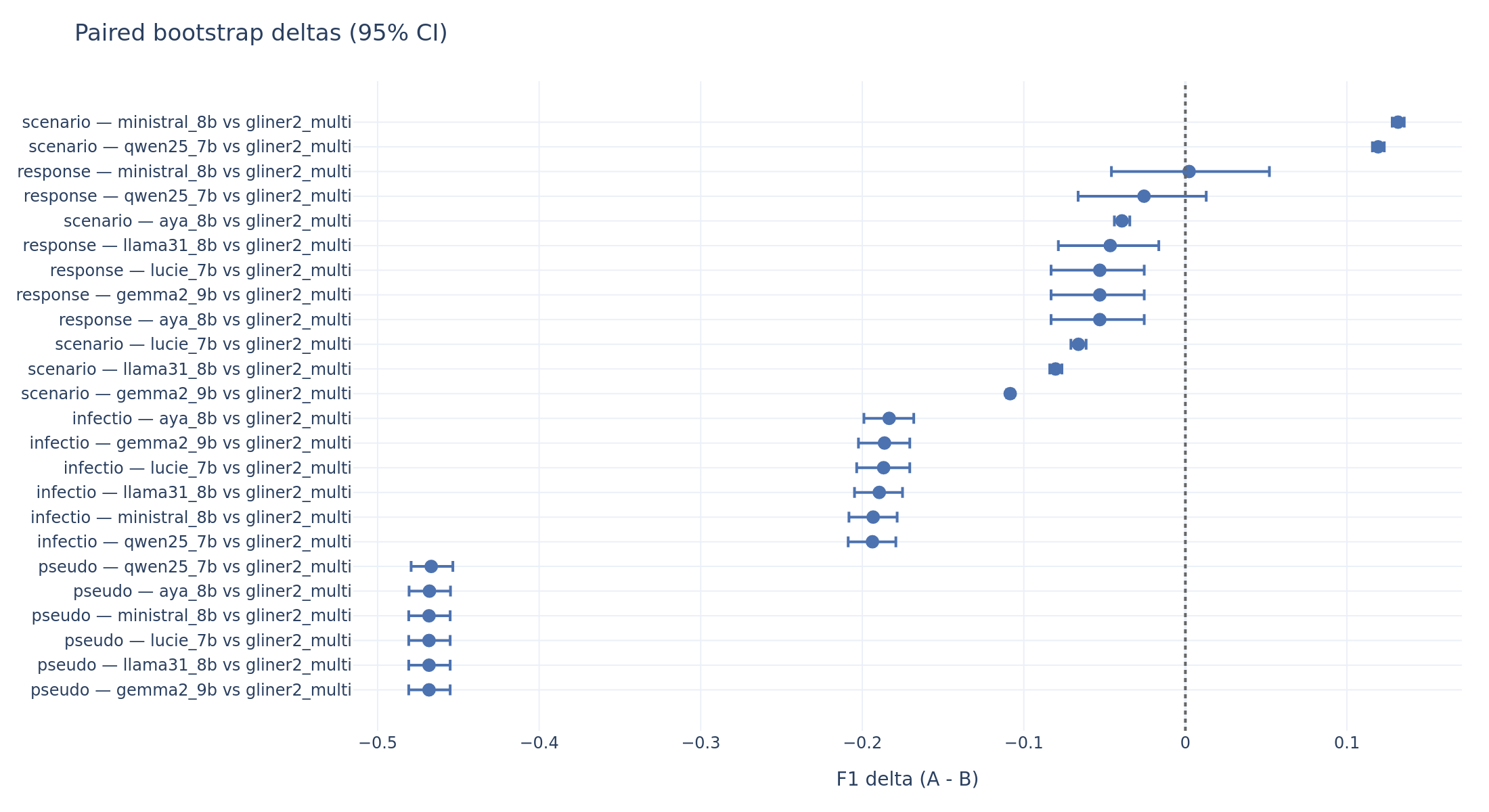
Paired bootstrap deltas (LLM minus GLiNER2, zero-shot). Negative values mean the LLM is worse. A CI that does not cross zero indicates statistical significance. Pseudonymization: every LLM is ~0.47 F1 below GLiNER2. Scenario: Ministral and Qwen are significantly above.
When Examples Help (and When They Don’t)
The few-shot track adds three fixed demonstration examples to the prompt. Everything else (seed, temperature, parser, gold corpus) is held constant, so the paired bootstrap on the delta between few-shot and zero-shot F1 isolates the effect of the demonstrations.
Three distinct patterns emerge.
Pattern 1: Recovery from collapse (Gemma2-9B). Gemma2 zero-shot scores effectively zero on scenario (F1 = 0.000) because it produces 0% schema-conforming outputs. Its few-shot scenario F1 jumps to 0.466, the highest of any system on any task. The +0.466 delta is not “learning from examples” in any meaningful sense. The demonstrations teach the model to produce valid JSON. This is recovery from an output format failure.
Pattern 2: Genuine improvement (Ministral, Qwen). Ministral gains +0.127 on scenario (from 0.240 to 0.367) and +0.148 on infectiology (from 0.002 to 0.149). Qwen gains +0.113 on infectiology and +0.107 on response. The confidence intervals exclude zero in every case. These models were already producing valid outputs on zero-shot; the demonstrations refine their extraction patterns.
Pattern 3: No effect (Llama, Lucie, Aya). Llama and Lucie show deltas near zero or slightly negative on most tasks. Lucie actually regresses on scenario (-0.021). Their schema conformity does not improve with demonstrations, suggesting these models do not incorporate the examples into their generation strategy.
Pseudonymization remains unsolved by few-shot prompting. Every model’s delta on pseudo is within the noise band of zero. The offset problem cannot be taught by three examples.
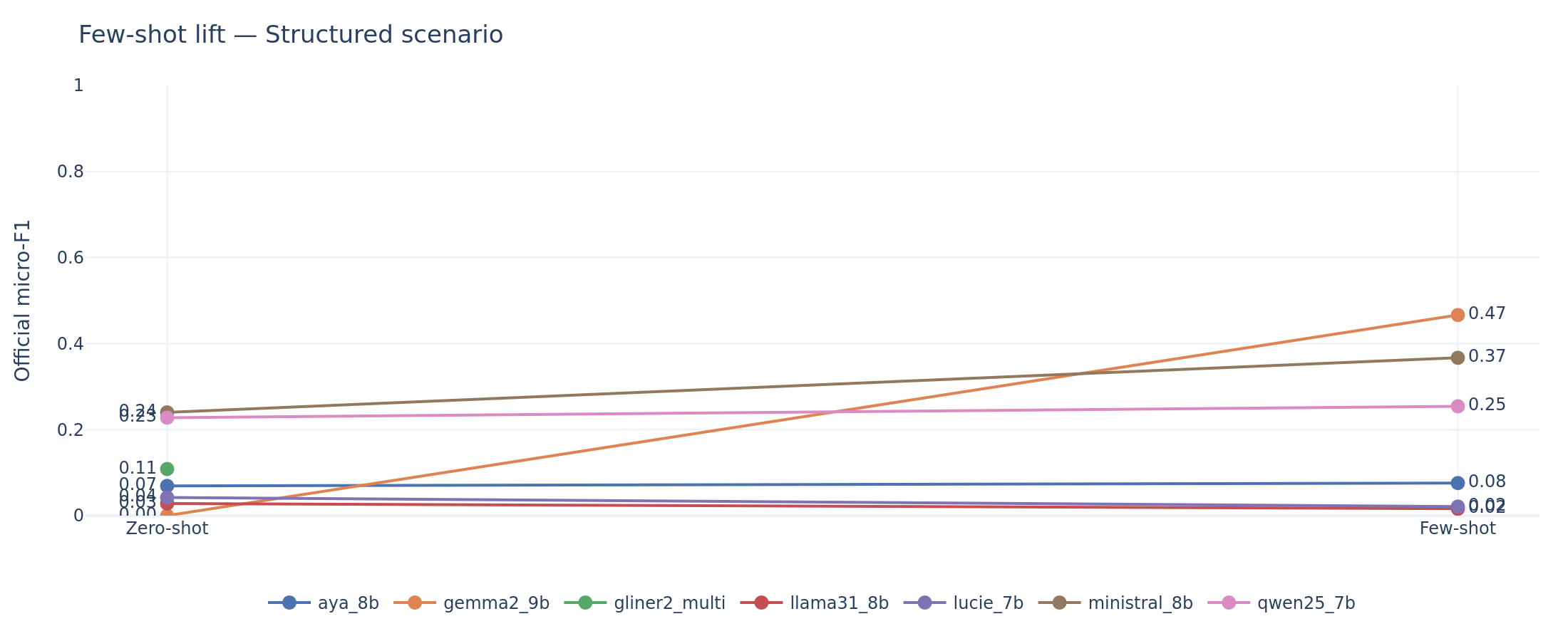
Scenario extraction: zero-shot to few-shot lift. Gemma2’s vertical jump from zero to 0.47 reflects format recovery, not learning. Ministral’s steady climb from 0.24 to 0.37 reflects genuine extraction improvement.

Infectiology: Ministral and Qwen show statistically significant gains from demonstrations. Lucie, Llama, and Aya remain flat.
For a clinician asking “if I write better prompts, will it work?”: the answer depends on both the task and the model. Few-shot helps the best-performing LLMs (Ministral, Qwen, Gemma2) on tasks where they already produce valid outputs. It does not solve fundamental architectural limitations (character offsets) or rescue models with poor output discipline.
The Real Bottleneck Is Not Clinical Understanding
Output Discipline as the Binding Constraint
This is the central analytical finding of the benchmark. The gap between what these models understand about clinical text and what they can reliably output as structured JSON is the dominant source of performance variance.
Consider Gemma2-9B on zero-shot. Its schema conformity is 0% on response and scenario, and 1.8% on pseudonymization. The model does not produce a single valid JSON envelope for response or scenario documents. Yet when it does produce valid output (in few-shot, where conformity rises to 58-68%), its F1 on scenario reaches 0.466, the best score in the entire benchmark. The clinical understanding is there. The generation discipline is not.
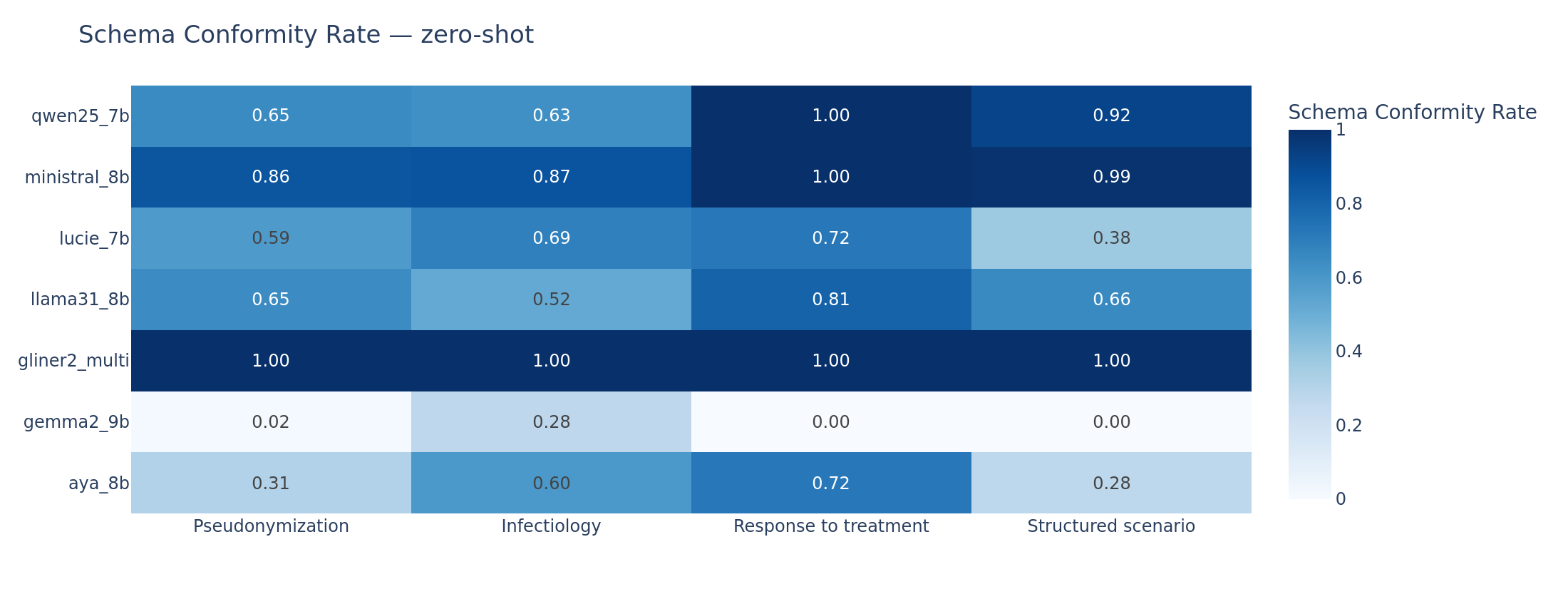
Schema conformity rate (proportion of documents where the model produces a valid JSON output matching the expected schema). Gemma2 zero-shot collapses across all tasks. Ministral maintains above 85% conformity throughout. GLiNER2 is at 100% by design.

Empty output rate (proportion of documents where the model produces no extractable records). The mirror image of schema conformity: Gemma2 zero-shot returns empty outputs for 100% of scenario documents.
A linear regression of per-task F1 on schema conformity, empty output rate, and token budget confirms that schema conformity is the single most explanatory variable of performance spread across models. The practical implication is that improvements in output-format reliability (via better constrained decoding, format-aware fine-tuning, or prompt engineering) would yield larger F1 gains than improvements in (proxies of) clinical understanding.
Key finding: The dominant bottleneck for SLMs on clinical IE is not medical knowledge but output discipline. Schema conformity explains more F1 variance than any clinical understanding metric.
Error Taxonomy: Two Failure Families
We classify every document-level error into categories based on what went wrong. Two families account for the vast majority of failures:
Invalid JSON errors occur when the model cannot produce a parseable JSON envelope. Gemma2 zero-shot generates ~4,960 invalid JSON errors. The model’s autoregressive decoding breaks down when faced with the structural constraints of the expected output format. This is a generation discipline problem that constrained decoding should, in principle, address (vLLM’s xgrammar backend was active, but it enforces JSON syntax, not semantic validity).
Offset drift errors occur when the model produces syntactically valid JSON with plausible-looking content, but the character offsets are wrong. This metric is tracked on pseudonymization where positions matters. Aya zero-shot generates ~3,440 offset drift errors. The model “knows” the entities (the text fields are often correct) but cannot compute or copy the right byte positions. This is an architectural limitation: autoregressive models do not have a native mechanism for counting characters in the input.
Ministral (133 invalid JSON, 1 offset drift on zero-shot) and Qwen (555 invalid JSON, 2 offset drift) are the cleanest models in output discipline, consistent with their leading LLM scores.

Error taxonomy (zero-shot). Two dominant families: Gemma2 is dominated by invalid JSON (generation discipline failure); Aya is dominated by offset drift (positional accuracy failure). Ministral and Qwen produce the fewest errors overall.
The Efficiency Question
Clinical pipelines process thousands of documents. Latency matters.
GLiNER2 is the dominant efficient system: ~1 second median latency per document with the best zero-shot global F1. Among LLMs, Ministral sits on the Pareto front at ~13-20 seconds median latency per document. Gemma2 few-shot matches Ministral’s few-shot global F1 but at roughly double the latency (~22-38 seconds depending on task).
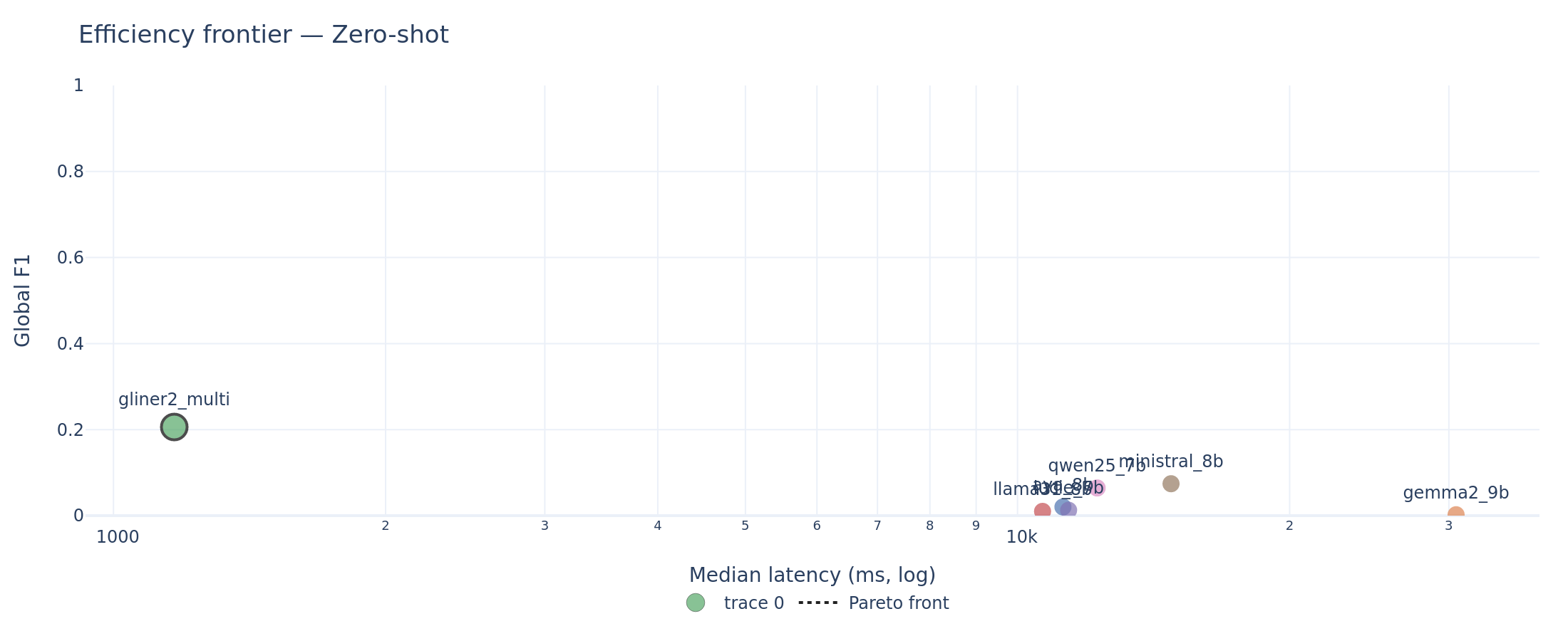
Efficiency frontier (zero-shot). GLiNER2 occupies the upper-left corner (high F1, low latency). The Pareto front connects GLiNER2 to Ministral. All other LLMs are dominated.
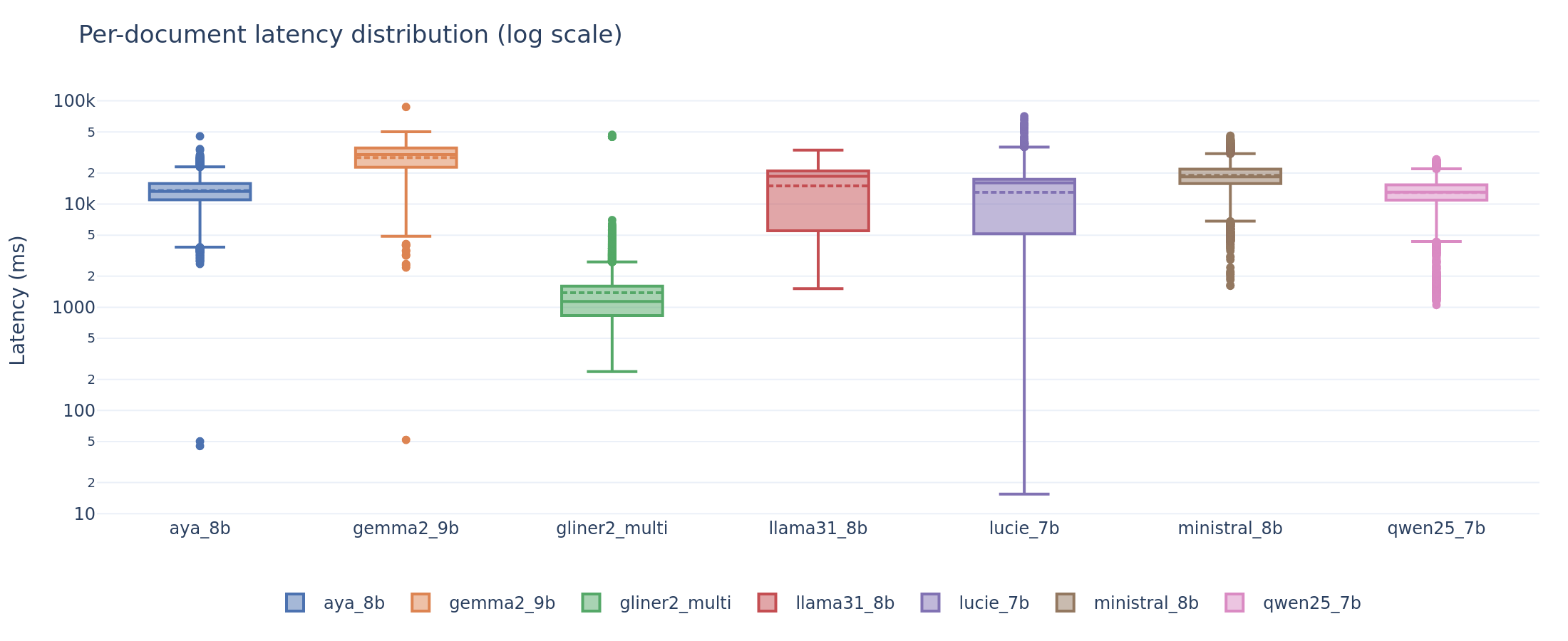
Per-document latency distribution. GLiNER2 completes in ~1 second. LLMs range from 5 to 38 seconds at the median, with significant variance across tasks and document lengths.
For a hospital pipeline processing a backlog of discharge letters, GLiNER2 processes documents 10-15x faster than the best LLM while achieving higher accuracy on three of four tasks. For structured scenario extraction, where LLMs are superior, Ministral-8B is the rational choice: it matches Gemma2’s few-shot F1 at a fraction of Gemma2’s latency.
Where Models Break Down
Aggregate F1 hides systematic weaknesses. We re-score each task on meaningful document slices: document-length quartiles, medical specialty, entity label, and (for infectiology) negation polarity.
Document length degrades pseudonymization. Already near zero for LLMs, pseudo F1 drops further on the longest document quartile. Longer documents contain more entities and more opportunity for offset drift. Even GLiNER2’s F1 decreases on the longest documents.
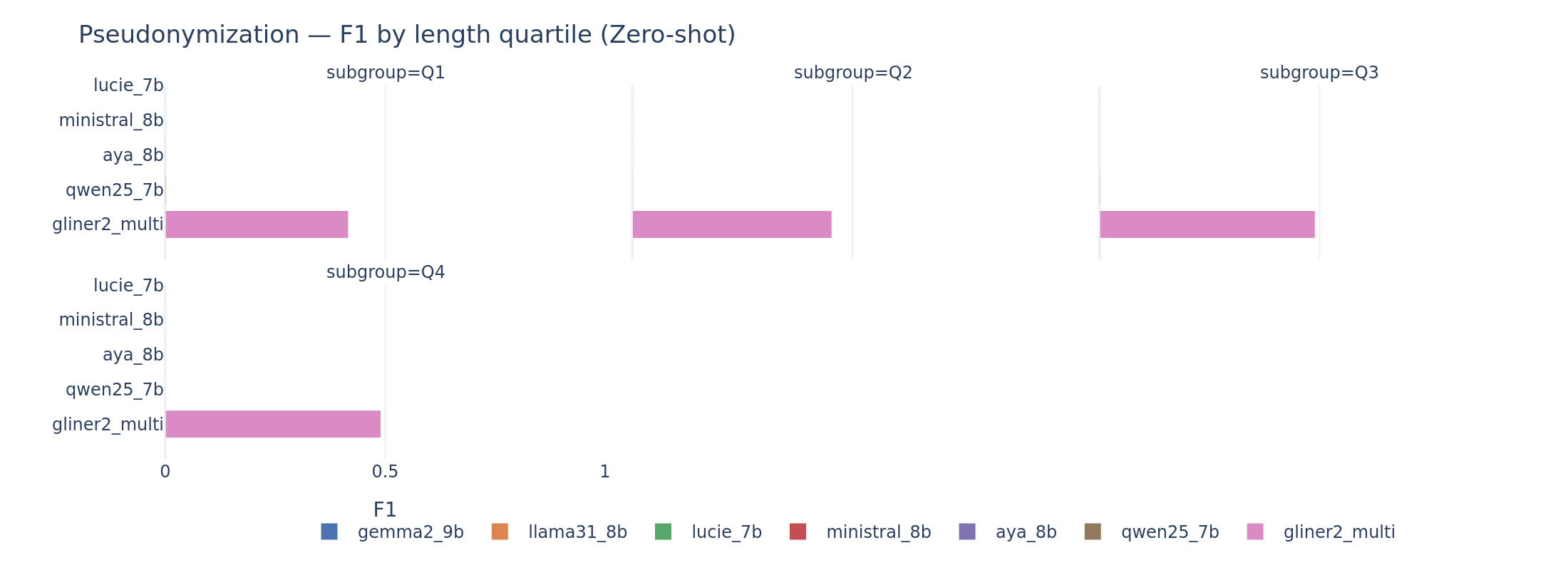
Pseudonymization by document length. Performance degrades on the longest quartile for all systems.
Negated mentions are the hardest for infectiology. The “Absent” polarity (negated infections, e.g., “absence de bacteriemie”) is systematically harder than “Present.” Models either miss the negation attribute entirely or fail to detect the mention when it appears in a negated context. French negation patterns (“pas de,” “absence de,” “aucun”) are handled inconsistently.
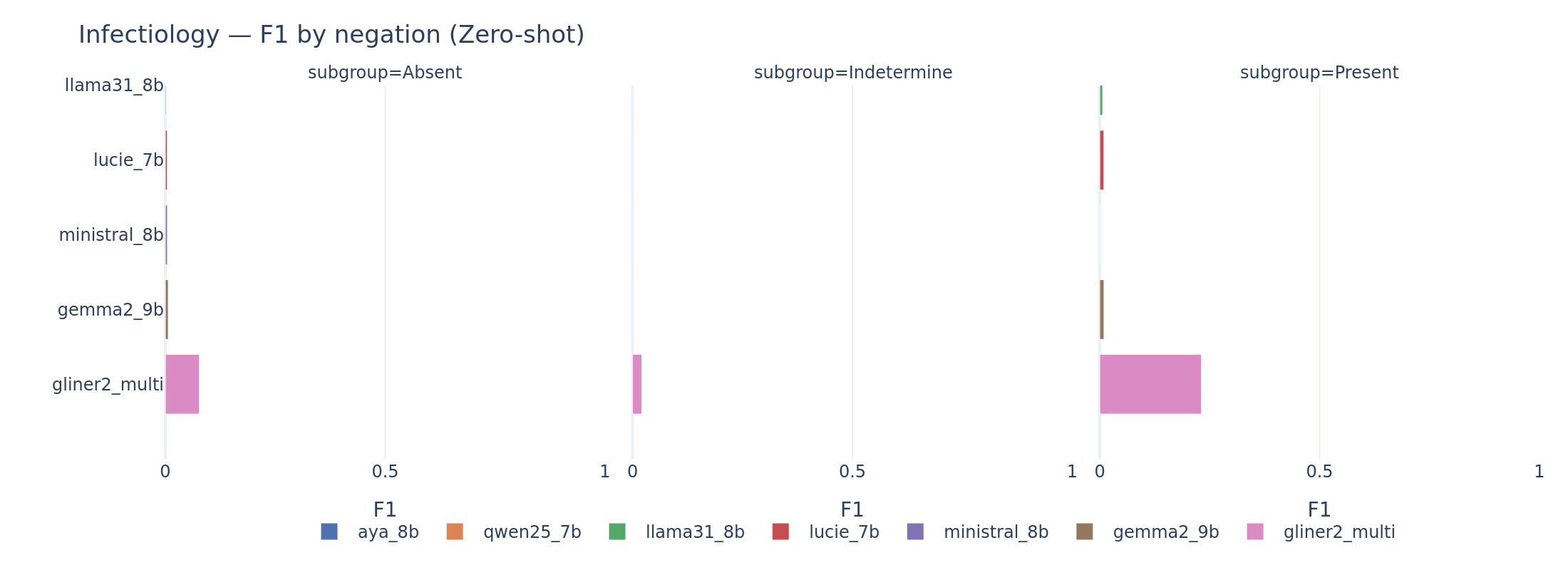
Infectiology by negation polarity. The “Absent” class is the hardest across all models. Negation detection in clinical French remains a significant challenge.
Performance varies by medical specialty on scenario. Cardiology notes, which tend to follow structured templates, produce higher F1 across models. Infectious disease notes, which are longer and more jargon-dense, produce lower F1. This specialty-level variance is practically important: a system that works well for cardiology may fail for infectious diseases.
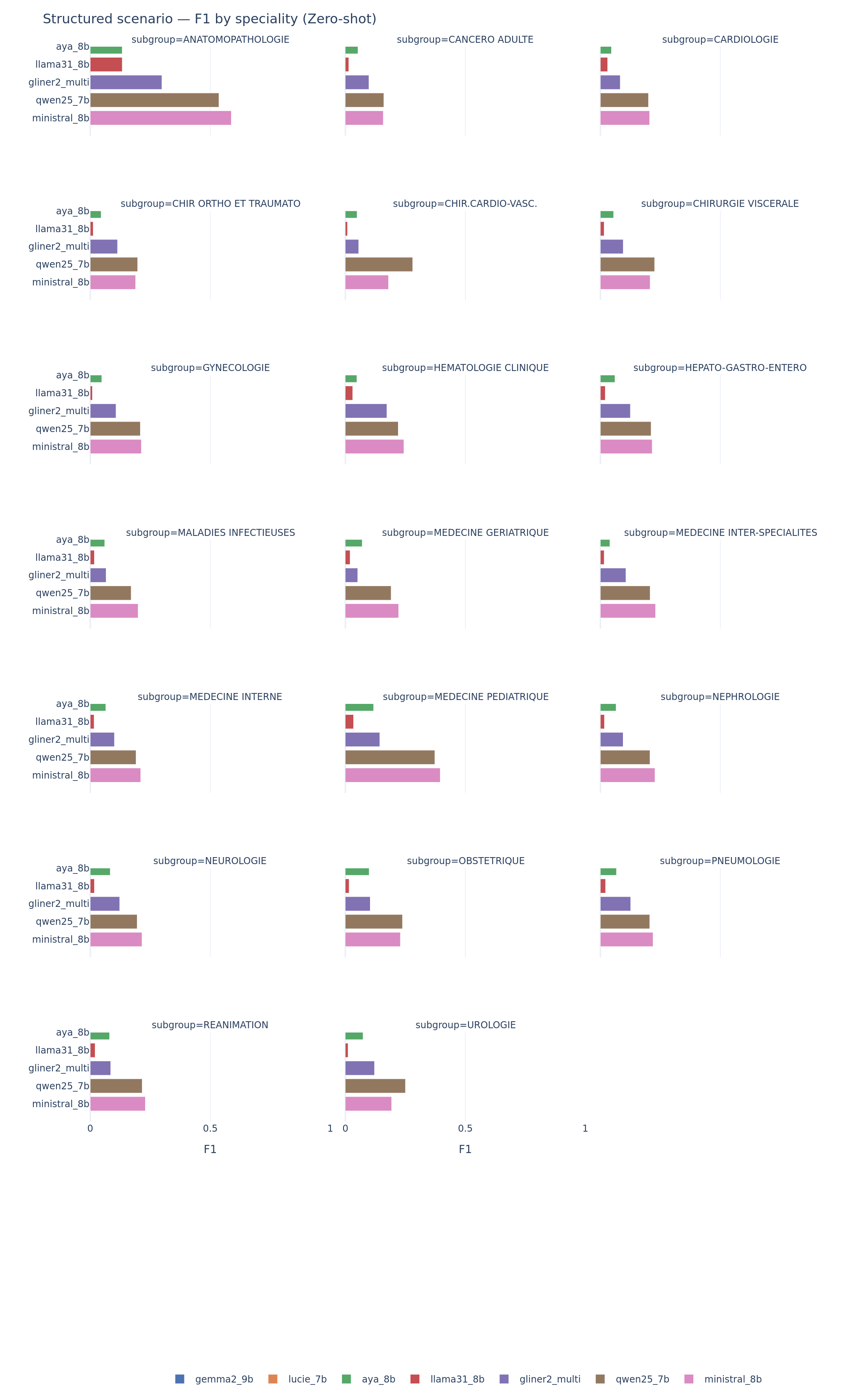
Scenario extraction by medical specialty. Performance varies strongly: models do well on structured specialties (cardiology) and struggle with long, jargon-dense notes (infectious diseases).
For clinicians: A global F1 number does not tell you whether the system will work for your specialty, your document length, or your specific extraction need. Always evaluate on data that matches your deployment context.
Discussion: What This Means for Clinical Deployment
The Readiness Gap
No system in this benchmark meets the performance thresholds that regulatory frameworks and clinical evaluation campaigns associate with safe deployment. The best single-task result, Gemma2 few-shot on scenario at F1 = 0.466, falls in what we define as the pilot zone (F1 between 0.40 and 0.70): viable as a pre-annotation tool with mandatory human review, but not as an autonomous system.
For pseudonymization, a safety-critical task under GDPR and health data regulations, even GLiNER2 at F1 = 0.468 is not deployable without human review. The i2b2 de-identification literature has established that systems below F1 ~0.95 leak too many identifiers for unsupervised use. The gap between current SLM performance and that threshold is enormous.
The distance between “interesting research result” and “deployable clinical tool” remains wide for zero-shot and few-shot approaches.
Encoder vs. Decoder: Architectural Fit Matters More Than Scale
The dominant variable in this benchmark is not parameter count but architectural match to the task type.
Span-labeling tasks favor encoders by design. Pseudonymization and infectiology require extracting text spans with precise boundaries. Encoder models like GLiNER2 process the input bidirectionally and score candidate spans directly. Autoregressive decoders must generate the span boundaries as text tokens, an inherently harder problem that introduces positional errors.
Structured extraction tasks favor decoders. The scenario task asks for high-level patient attributes (age, diagnosis, admission mode) from free text. This is closer to reading comprehension than to sequence labeling. Decoders’ generalist language understanding gives them an advantage over encoders that are optimized for local span detection.
This observation generalizes beyond this specific benchmark. It is a structural finding about how different architectures handle different task types. The implication for practitioners: choose the architecture that matches the task, not the one with the most parameters.
Output Discipline as a First-Order Concern
Across all models and tasks, schema conformity explains more F1 variance than any measure of clinical text understanding. This finding has implications beyond the models tested here.
The practical bottleneck for clinical AI deployment is often not “does the model understand the clinical text” but “can it reliably produce a valid structured output.” A model that extracts correct information 60% of the time but produces invalid JSON 40% of the time is not a 0.60-F1 system. It is a broken pipeline. Robustness metrics (schema conformity, output validity, empty output rate) should be first-class citizens in any clinical NLP evaluation, not afterthoughts.
The Path Forward
Several directions could narrow the gap:
- Supervised fine-tuning on PARHAF or similar clinical corpora is the natural next step. Even lightweight LoRA adaptation could dramatically improve output discipline and task-specific extraction.
- Hybrid pipelines: use GLiNER2 (or a similar encoder) for span-labeling tasks and an LLM for structured extraction. The two architectures are complementary, not competing.
- Advanced prompting strategies: chain-of-thought for offset computation, self-consistency for robust extraction, or tool-augmented approaches where the model calls a character-counting utility.
- Larger models: systems above 9B parameters, or reasoning-specialized models, may close the gap, though at higher computational cost.
Caveats and Limitations
A handful of caveats bound the interpretation of these results.
Single-seed inference. Every score is based on a single deterministic run (temperature 0, seed 42). The bootstrap confidence intervals capture sensitivity to document resampling, not to decoding randomness. A cross-seed stability study would strengthen the conclusions.
Fixed few-shot bank. We cannot separate “the model benefits from demonstrations in general” from “the model benefits from these specific three examples.” A sensitivity study over alternative demonstration banks is left for future work.
Strict offset metric. The pseudonymization metric requires exact (start, end) character matches. A relaxed overlap metric (e.g., token-level or partial-span credit) would give a more generous picture of LLM capabilities on entity detection. The strict metric is appropriate for evaluating deployment readiness (a span off by one character leaks PHI), but it obscures the models’ genuine entity recognition ability.
Language and corpus scope. All datasets are in French from a single institutional source (Health Data Hub). While the meta-level findings of this benchmark (architectural fit, output discipline patterns, few-shot behavior) are language-agnostic, the absolute F1 numbers may not transfer directly to English or other languages where models have seen more clinical training data. Replication on English clinical corpora (e.g., i2b2, n2c2) would strengthen generalizability claims.
No fine-tuned models. This is a zero-shot and few-shot benchmark. Supervised fine-tuning would likely change the picture substantially, particularly for output discipline.
No advanced prompting. We evaluate a single prompting format (instruction + schema + document, optionally with demonstrations). Chain-of-thought, self-consistency, retrieval augmentation, and tool use are not tested.
Readiness thresholds. The 0.40 and 0.70 thresholds we use as reference points are pragmatic, informed by regulatory expectations and prior evaluation campaigns, but they are not universal standards. The appropriate threshold depends on the specific deployment context and risk profile.
Practical Recommendations
For clinicians and hospital IT teams
-
Do not deploy any tested system for autonomous pseudonymization without any further tuning. Even GLiNER2, the best performer at F1 = 0.47, misses too many identifiers for unsupervised use. Use it as a pre-annotator with mandatory human review. More precise parameter tuning, such as adjusting GLiNER’s cosine similarity threshold in favor of a better recall, and/or targeted model fine-tuning could help improve robustness.
-
For structured scenario extraction, Ministral-8B few-shot is the strongest SLM option (F1 = 0.37). Plan for human-in-the-loop validation. Gemma2-9B few-shot reaches higher F1 (0.47) but at double the latency and with lower robustness.
-
Budget 10-15x more compute for LLM-based pipelines compared to encoder-based ones. GLiNER2 processes documents at ~1 second; LLMs require 13-38 seconds (including time to first token delay).
For AI engineers / scientists
-
Offset-aware decoding is an open research problem. Autoregressive models fundamentally struggle with character-level positional constraints. This is not a prompting problem but an architectural one. Solutions might involve pointer networks, copy mechanisms, or hybrid architectures that combine span scoring with text generation.
-
Report robustness metrics alongside accuracy in clinical NLP evaluations. A model that produces correct extractions 40% of the time and invalid outputs 60% of the time is a broken pipeline, not a 0.40-F1 system. Schema conformity, empty output rate, and JSON validity should be standard columns in every benchmark table.
-
Replication on English clinical corpora (i2b2, n2c2) would test whether the meta-level findings of this study hold cross-linguistically. We expect the architectural-fit and output-discipline observations to generalize; the absolute F1 numbers will differ.
Reproducibility
Every result in this report is reproducible from the open-source codebase:
- Code and configuration: github.com/lounesmechouek/parhaf-clibench
- Evaluation suite:
configs/suites/v1_full.yamldefines the exact experimental setup (models, tasks, tracks, parameters) - Prompt templates: Jinja2 templates in
prompts/, versioned by SHA-256 hash - Dataset provenance: All four PARHAF subsets are loaded from HuggingFace with pinned Git revisions, cached locally with a stable fingerprint (SHA-256 over dataset name, example count, document IDs, and text hashes)
- Inference parameters: temperature = 0, top-p = 1, seed = 42, max tokens = 512 (all LLMs)
- Bootstrap: B = 1,000 document-level replications, seed = 42, percentile 95% CI
- Scoring audit: 52/52 cells reproduce shipped values to six decimal places
- Hardware: NVIDIA RTX A6000, single GPU
The full prediction set (65,065 documents) and all intermediate artifacts (scores, robustness metrics, error logs, timing data) are preserved in the run directory.
Conclusion
This benchmark set out to answer a practical question: can 7-9B parameter language models reliably extract structured clinical information from discharge letters? The answer, after 65,065 predictions on real physician-written notes, is: not yet, but the reasons are more informative than the headline.
The gap between SLM capabilities and clinical deployment readiness is not primarily about medical knowledge. These models often understand the clinical text well enough to identify the right entities, classify the right outcomes, and extract the right fields. The bottleneck is output discipline: producing valid, schema-conforming structured outputs with correct positional information. That is an engineering problem, not a fundamental capability limit.
The benchmark also reveals that architectural fit matters more than parameter count. A 200M-parameter encoder outperforms every 7-9B decoder on span-labeling tasks, while the decoders win on structured extraction. The right tool for the job depends on the job.
PARHAF, as the first large-scale corpus of realistic physician-written clinical notes with multi-task annotations, provides valuable infrastructure for this kind of rigorous evaluation. We hope this benchmark serves as a baseline that future work on fine-tuning, hybrid architectures, and advanced prompting will improve upon, and that replication on other languages and clinical corpora will test the generality of these findings.
The gap between SLM hype and clinical readiness is real but measurable. Measurable problems are solvable problems. The code and data references open-source.
Benchmark run: April 2026. Models evaluated with weights available as of April 2026. PARHAF dataset version: March 2026 release.